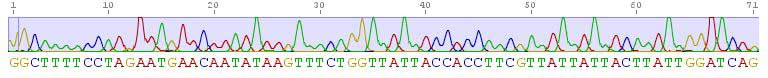
DNA barcodes

DNA barcoding
“DNA barcoding” is a new term that means using DNA sequence data to help identify species. The main initiative for large-scale use of DNA for identification has come from Canada (Canadian Barcode of Life Network). An international initiative (CBOL) has now been launched to barcode all living organisms (see links at left).
Barcodes could be particularly useful for microbes, including diatoms, because of the difficulty we have in telling one species from another under the microscope. For animals and red algae a short section (about 700 bp long) of a mitochondrial gene, cytochrome oxidase 1 (or cox1), has been found to be a useful tool to discriminate one species from another. This region is now being tested for its suitability in microscopic organisms. Curiously, in green plants (mosses, liverworts, ferns, flowering plants, green algae), cox1 appears to be useless, because it is not variable enough.
For a number of years before the recent DNA barcoding push, we were using a section of a chloroplast gene to distinguish one diatom species from another; this was the gene (rbcL) coding for the large subunit of RuBisCO (ribulose-1,5-bisphosphate carboxylase/oxygenase), which is the primary carboxylating enzyme involved in photosynthesis. With the advent of DNA barcoding, we decided to assess the usefulness of cox1 for distinguishing diatom species, using the S. pupula species complex as a test (Evans et al. 2007). We compared the performance (in terms of the amount of DNA variation within species compared to that between species) of cox1 to rbcL and to two nuclear sequences from the ribosomal RNA cistron in the 18S (small-subunit) and ITS (internal transcribed spacer) regions. We found cox1 to be more variable than rbcL and 18S and much more convenient to use than ITS (because we didn’t need to clone PCR products and alignment of ITS can be problematic) and so we supported the use of cox1 as an additional tool in diatom identification. This was the first study to investigate a potential barcoding region for diatoms.
During the past year or more, we have routinely used cox1 barcodes to assess Sellaphora diversity and identify newly isolated clones. The partial gene sequences also allow us to make a preliminary estimate of evolutionary relationships, which can then be tested further by using other genetic data.
Reference
Evans, K.M., Wortley, A.H. & Mann, D.G. (2007). An assessment of potential diatom “barcode” genes (cox1, rbcL, 18S and ITS rDNA) and their effectiveness in determining relationships in Sellaphora (Bacillariophyta). Protist 158: 349–364.

 This site is hosted by the Royal Botanic
Garden Edinburgh.
This site is hosted by the Royal Botanic
Garden Edinburgh.